News
MAGNet Releases Web Browser-Enabled Version of geWorkbench Bioinformatics Software Platform
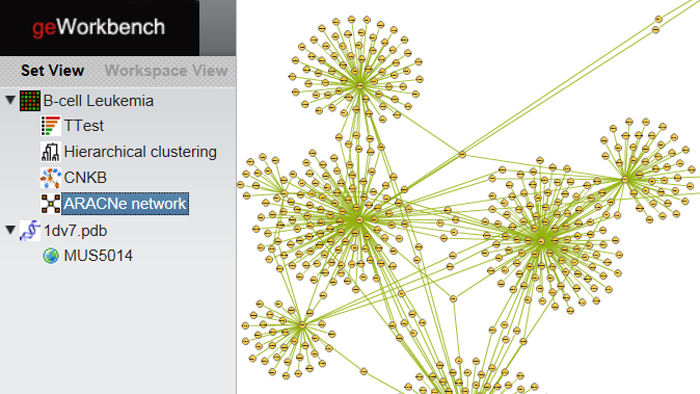
A new version of geWorkbench lets researchers access a range of powerful, integrated bioinformatics tools using a standard web browser. Here, an ARACNe-generated gene regulatory network is displayed using the Cytoscape Web plugin.
Since its creation in 2005, investigators in Columbia University’s Center for the Multiscale Analysis of Genomic and Cellular Networks (MAGNet) have developed a large number of computational tools for studying biological systems from the perspectives of structural biology and systems biology. To consolidate and disseminate these tools to the wider research community, MAGNet developed geWorkbench (genomics Workbench), a free, open-source bioinformatics application that gathers all of the Center’s software and databases into one integrated software platform. These include applications for the analysis of cellular regulatory networks, protein structure, DNA and protein sequences, gene expression, and other kinds of biological data.
Initially, geWorkbench was made available as a software package that users could install and run on their local computers. Now, in a major upgrade, MAGNet has released a web-based version that makes these tools accessible through a browser interface.
Aris Floratos, an assistant professor in the Columbia University Department of Systems Biology and the leader of the development team behind geWorkbench, says that removing the burden of software installation will make it easier for users to access the wealth of computational research resources that MAGNet offers. “Many of the tools created at MAGNet labs are still under active development, and in some cases new versions are released every few months," Dr. Floratos explains. "In the past, access to new versions required reinstallation of the application. Having a web interface eliminates this overhead. Once we make updates to our server, they are available to the community immediately.”
An important part of MAGNet’s vision for systems biology is that researchers today need to look simultaneously at multiple types of biological data in order to understand how an entire system functions. From its beginnings, geWorkbench was designed to facilitate this integrative approach. Using a collection of visually linked interfaces, researchers can browse, manipulate, and visualize data in a variety of ways. They can design workflows that move data from one component to the next, and can access tutorials that both explain the algorithms that drive the software and provide instructions for each individual software module. Floratos stresses the importance that his team has placed on making geWorkbench user friendly, saying, “A big focus of what we’re doing is to make the tools accessible to both computational and non-computational biologists. This will enable the widest possible community of researchers to benefit from the tools that MAGNet researchers have developed.”
In addition to including tools developed by MAGNet, geWorkbench provides access to third-party components that expand its capabilities. The platform’s developers have been working with researchers at the Broad Institute to connect geWorkbench to GenePattern, a complementary genomic analysis platform that provides additional tools for gene expression analysis, proteomics, SNP analysis, flow cytometry, RNA-seq analysis, and other data processing tasks. Researchers can now take data from geWorkbench, move it into GenePattern for analysis, and port the results back into geWorkbench for additional calculation. GeWorkbench enables similar access to other resources including the Protein Data Bank, Gene Ontology, and BLAST queries at the National Center for Biotechnology Information (NCBI).
Other resources that have recently become available in geWorkbench include access to the Cellular Networks Knowledge Base, a public database of interaction networks, and a new database of drug synergy measurements developed by the laboratory of Andrea Califano in conjunction with the Library of Integrated Network-Based Cellular Signatures (LINCS) at the National Institutes of Health (NIH).
GeWorkbench also provides access to GenSpace, an innovative framework that enables collaboration and interaction among geWorkbench users. Developed in the laboratory of Gail Kaiser, a MAGNet member and professor in the Columbia University Department of Computer Science, GenSpace logs users’ workflows to provide information about how they interact with geWorkbench. This makes it possible to provide recommendations to users about the most efficient workflows for performing certain tasks. GenSpace also includes a social networking component that enables users to create circles of friends, communicate with others, and gain advice from expert users of different tools. In this way, GenSpace leverages community knowledge to improve users’ experience of the tools and the application.
The current release of the web-based geWorkbench is a public beta version. Dr. Floratos anticipates that it will grow and eventually completely replace the downloadable desktop version. “A number of the marquee MAGNet tools have already been ported into the web version and more are coming soon,” he points out. “We welcome people to use it and provide us with feedback.”
— Chris Williams