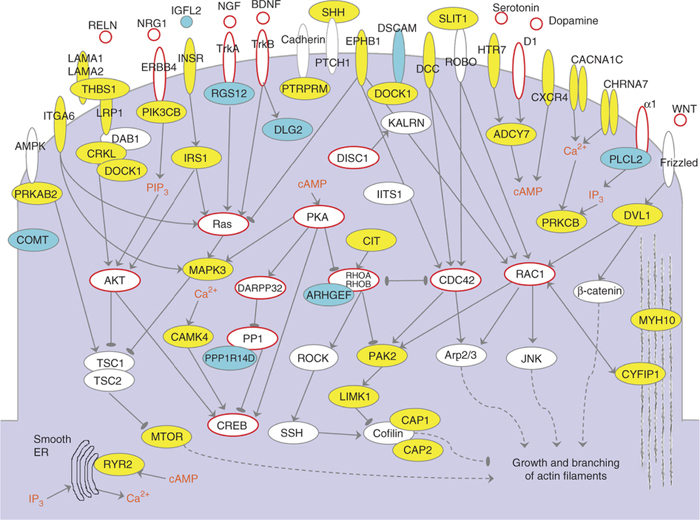
Genes forming cluster I in the context of cellular signaling pathways. Proteins encoded by cluster genes are shown in yellow, and those corresponding to other relevant genes that were present in the input data but not selected by the NETBAG+ algorithm are shown in cyan.
In a new paper published in the journal Nature Neuroscience, Columbia University researchers report that many of the genes that are mutated in schizophrenia are organized into two main networks. Surprisingly, the study also found that a genetic network that leads to schizophrenia is very similar to a network that has been linked to autism.
Using a computational approach called NETBAG+, Dennis Vitkup and colleagues performed network-based analyses of rare de novo mutations to map the gene networks that lead to schizophrenia. When they compared one schizophrenia network to an autism network described in a study he published last year, they discovered that different copy number variants in the same genes can lead to either schizophrenia or autism. The overlapping genes are important for processes such as axon guidance, synapse function, and cell migration — processes within the brain that have been shown to play a role in the development of these two diseases. These gene networks are particularly active during prenatal development, suggesting that the foundations for schizophrenia and autism are laid very early in life.